What are data federation and federated analysis?
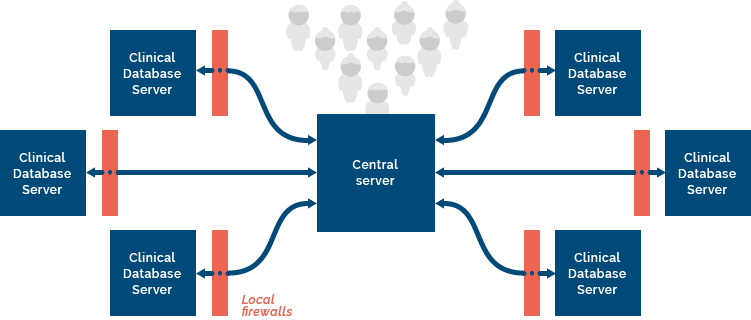
Data federation simply means connecting databases so they can talk to each other. In RHAPSODY we have built a federated infrastructure based on open source technology that enables powerful statistical analysis to be performed on patient data hosted on servers (federated nodes) at different locations across Europe.
Why do we need a federated infrastructure?
There are often legal or ethical constraints that prevent the storing of individual patient-level data on a central repository for analysis, or the transferring of data between participating centres in a research project. The federated infrastructure we have put in place protects the patient data so that it never leaves the server on which it is hosted, whilst at the same time enabling analysis to be performed remotely on each server. We describe this analysis is ‘non-disclosing’, meaning that analysis is performed without any sensitive patient data being accessed.
How many cohorts are in the RHAPSODY Federated Database?
So far, we have federated 12 patient cohorts totaling over 68000 individuals (see Table). The work involved painstaking reformatting of the data from each cohort to a common standard (CDISC) and harmonisation of the data using standard ontologies and measurement units. Once completed, each harmonised dataset was loaded into its respective node and made available for federated analysis.
What different kinds of patients cohorts are available in the Federated Database?
Several different types of patient cohorts are available in the database (see Table). There are 6 cohorts of pre-diabetic individuals, these are people who show signs of impaired glucose tolerance but have not been diagnosed with type 2 diabetes; 4 cohorts of diabetes progression, where all individuals have been diagnosed with type 2 diabetes and where the progression of the disease in each individual is followed over time; 1 gastric bypass cohort where individuals with type 2 diabetes who have undergone gastric bypass surgery are followed over time to measure how their diabetes develops; and one clinical trial where response of patients to therapeutic intervention for "lipid lowering" was measured in a randomized, controlled study.
How does the RHAPSODY Federated Database work?
Data analysis through the federated database is made possible through the R statistical programming environment. A user logs on to the database from within the R environment running on their local computer (e.g. using RStudio). He or she can then use various data mining and statistical tools that are available through the system. We have developed a federated analysis package called dsSwissKnife (publicly available on GitHub) that provides these tools. Further information about the tools and how they work can be found here.
What is the future of the Federated Database beyond RHAPSODY?
The RHAPSODY federated database is a key asset, enabling research to fight the global epidemic of type 2 diabetes. Therefore, we have worked on ensuring that the federated database will be available and maintained after the end of the project, with highly regulated access procedures in place. Part of this work has involved creating a public website where information can be found on which patient cohorts are included and how to apply for access to use the system. In the future, if funding is available, we also plan to add new cohorts as well as periodically updating the data in existing cohorts. The number of statistical tools available to users of the system will also evolve over time, offering new ways to analyse the cohort data.
Clinical Data Interchange Standards Consortium. A set of widely used standards for clinical data.
A research study in which one or more human subjects are prospectively assigned to one or more interventions (which may include placebo or other control) to evaluate the effects of those interventions on health-related biomedical or behavioral outcomes (source: https://grants.nih.gov/).
Patient cohort: A group of individuals affected by common diseases, environmental or temporal influences, treatments, or other traits whose progress is assessed in a research study (source: https://medical-dictionary.thefreedictionary.com/)
Computer simulation models are a common tool to predict how different treatments may affect disease progression and outcomes. They are used to extrapolate the findings of clinical trials into lifetime costs and health benefits.
Diabetes is a chronic disease characterised by chronically elevated levels of blood sugar (hyperglycemia), due to alterations in the production or the use of insulin by the body. Insulin action is vital to ensure that glucose (sugar, basic source of energy) from our food is properly utilised by cells of the human body.
Insulin is a hormone produced by the pancreas (more precisely by pancreatic beta cells), which is crucial to control blood glucose (glycemia). Once secreted into the blood stream, insulin orchestrates a coordinated response to lower blood glucose by acting on several tissues (namely insulin-target tissues). Notably, insulin stimulates glucose uptake by muscle and fat (adipose tissue) and prevents glucose production by the liver.
For a long time, diabetes has been labelled either ‘type 1’ or ‘type 2’ diabetes. Type 2 diabetes is the result of failures across several complex biological systems. Compared to those with type 1 diabetes, patients with type 2 diabetes can still produce insulin, at least during the first stages of the disease (prediabetes and early type 2 diabetes). However, their insulin-target tissues stop responding to the normal amounts of insulin produced after eating: this so-called “insulin resistance” process prevents glucose uptake by cells, resulting in rise of blood glucose. Chronic hyperglycemia tricks the pancreas into over-producing insulin. If untreated, this vicious circle of events can spiral out of control leading to an exhaustion of the pancreas (“pancreatic beta cell failure”) to produce insulin (insulin deficiency).
Diabetes is a major public health problem since over time, it can damage heart and blood vessels (heart attacks and strokes), nerves (neuropathy resulting in foot ulcers and possibly limb amputation), eyes (retinopathy leading to blindness) and kidneys (nephropathy).
Epigenetics is the study of how behaviors and environment can cause changes that affect the way genes work. Unlike genetic changes, epigenetic changes are reversible and do not change DNA sequence, but they can change how the body reads a DNA sequence (source: https://www.cdc.gov/genomics/disease/epigenetics.htm).
Elovl2: this gene encodes for the protein called "Elongation of very long chain fatty acids protein 2" which catalyzes the first and rate-limiting reaction of the four reactions that constitute the long-chain fatty acids elongation cycle. This endoplasmic reticulum-bound enzymatic process allows the addition of 2 carbons to the chain of long- and very long-chain fatty acids (VLCFAs) per cycle (source: https://www.uniprot.org/uniprot/Q9JLJ4)
A server containing a clinical database which can be connected to from a central computer using web services (HTTPS)
A surgical bypass operation that typically involves reducing the size of the stomach and reconnecting the smaller stomach to bypass the first portion of the small intestine so as to restrict food intake and reduce caloric absorption in cases of severe obesity (source: https://merriam-webster.com/).
Organs where insulin exerts its blood glucose lowering action. Liver: insulin decreases glucose production; muscles and adipose tissue: insulin stimulates glucose uptake.
Multi-omics refer to biological information in the form of any molecules in our bodies which are part of our everyday functioning and can change during diseases.
Genomics: study of the full genetics components of an organism (the genome) which includes the entire DNA sequence. It also considers the inter-relationships between the genes and their interaction with environmental factors.
Transcriptomics: transcriptomics is used to identify the qualitative and quantitative RNA levels in the whole genome. It can tell us which transcripts are present together with the levels of their expression. Almost 80% of the genome is transcribed to RNA.
Proteomics: study of the structure, function and physiological role of the entire proteins in a cell or tissue of an organism.
Metabolomics:study of the metabolites present in a cell or tissue/fluids of an organism. This includes small molecules, carbohydrates, peptides, lipids, nucleosides and metabolism products.
A set of concepts and categories in a subject area or domain that shows their properties and the relations between them.
A phenotypic trait is an obvious, observable, and measurable trait; it is the expression of genes in an observable way. An example of a phenotypic trait is a specific hair color. Underlying genes, which make up the genotype, determine the hair color, but the hair color observed is the phenotype. The phenotype is dependent on the genetic make-up of the organism, and also influenced by the environmental conditions (source: https://en.wikipedia.org/wiki/Phenotypic_trait)
PNLIPRP1: this gene encodes for the protein called "pancreatic lipase-related protein 1" which may function as inhibitor of dietary triglyceride digestion (source: https://www.uniprot.org/uniprot/P54315)
Precision medicine refers to the tailoring of medical treatment to the individual characteristics of each patient, […] the ability to classify individuals into subpopulations that differ in their susceptibility to a particular disease, in the biology or prognosis of those diseases they may develop, or in their response to a specific treatment (definition from the US National Research Council, https://en.wikipedia.org/wiki/Precision_medicine).
When fasting blood glucose is raised beyond normal levels, but is not high enough to warrant a diabetes diagnosis (source: https://diabetes.co.uk/)
Health benefits are often expressed as quality-adjusted life years (QALYs). QALYs combine longevity and quality of life in a single metric and allow comparisons across different treatments and diseases. As a result, the impact on QALYs are required by several European agencies to make decisions about whether to adopt and reimburse medicines at a national level.
A programming language and free software environment for statistical computing and graphics supported by the R Foundation for Statistical Computing. The R language is widely used among statisticians and data miners for developing statistical software and data analysis (source: https://en.wikipedia.org/wiki/R_(programming_language)).
An integrated development environment for R that runs on Windows, Mac or Linux operating systems.
Sensitivity is the proportion of true-positives which actually test positive, and how well a test is able to detect positive individuals in a population (source: https://www.fws.gov/aah/PDF/SandS.pdf).
Specificity is the proportion of true-negatives which actually test negative, and reflects how well an assay performs in a group of disease negative individuals (source: https://www.fws.gov/aah/PDF/SandS.pdf).




This work has been supported by the Swiss State Secretariat for Education‚ Research and Innovation (SERI) under contract number 16.0097-2.
The opinions expressed and arguments employed herein do not necessarily reflect the official views of these funding bodies.




